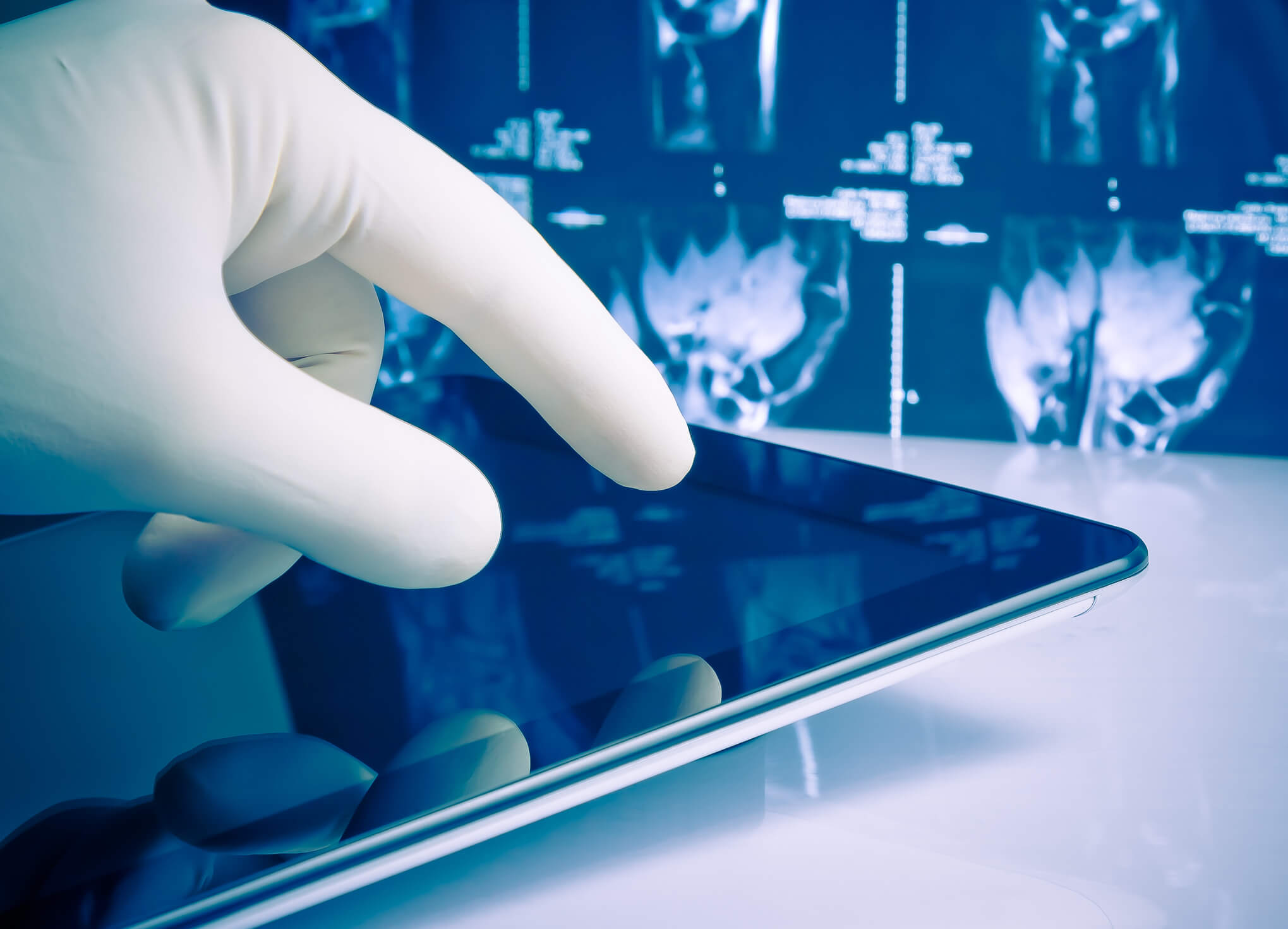
In order to naturally complement pathologists workflow to detect cancer cells, American multinational tech giant Google has applied Deep Learning, a class of machine learning algorithms, to digital pathology.
“We used images (provided by the Radboud University Medical Centre, the Netherlands) which have also been used for the 2016 ISBI Camelyon Challenge1 to train algorithms that were optimised for localisation of breast cancer that has spread (metastasised) to lymph nodes adjacent to the breast,” said Martin Stumpe, Technical Lead, and Lily Peng, Product Manager at Google in a blog post.

Stumpe and Peng said that for cancer a pathologists diagnosis has a profound impact on a patients therapy and the reviewing of pathology slides is a very complex task, requiring years of training to gain the expertise and experience to do well.

“Pathologists are responsible for reviewing all the biological tissues visible on a slide. However, there can be many slides per patient, each of which is 10+ gigapixels when digitised at 40X magnification. Imagine having to go through a thousand 10 megapixel (MP) photos, and having to be responsible for every pixel. Needless to say, this is a lot of data to cover, and often time is limited,” they added.

To address these issues, Stumpe and Peng applied deep learning to digital pathology with positive results.

“After additional customisation, including training networks to examine the image at different magnifications (much like what a pathologist does), we showed that it was possible to train a model that either matched or exceeded the performance of a pathologist who had unlimited time to examine the slides,” said the authors.
In fact, the prediction heatmaps produced by the algorithm had improved so much that the localisation score (FROC) for the algorithm reached 89 per cent, which significantly exceeded the score of 73 per cent for a pathologist with no time constraint.
Training models is just the first of many steps in translating interesting research to a real product. “But we are off to a very promising start, and we hope by sharing our work, we will be able to accelerate progress in this space,” Stumpe and Peng said.
Be a part of Elets Collaborative Initiatives. Join Us for Upcoming Events and explore business opportunities. Like us on Facebook , connect with us on LinkedIn and follow us on Twitter , Instagram.
"Exciting news! Elets technomedia is now on WhatsApp Channels Subscribe today by clicking the link and stay updated with the latest insights!" Click here!