Researchers use antibodies as basic tools in human and animal health fields such as research, diagnostics, experiments and treatment development. To comprehend, how normal cells function and how they vary from diseased cells, scientists sometimes utilize antibodies to target and identify specific proteins that are in the cells at a certain stage of growth and development. Using the results of such experiments, they can then create a prototype of how the cell works, and what happens when it becomes problematic and diseased. This is valuable information for developing and testing new treatments for the disease.
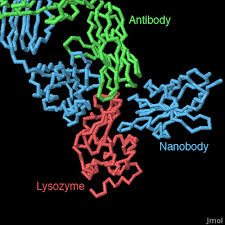
Nanobodies, which were first discovered several years ago in the unique antibodies of camels and llamas – offer an exciting alternative to antibodies as biomedical tools because they are much smaller, and show higher affinity to their molecular targets. Nanobodies have about one-tenth of the weight of antibodies, and they are stable and easy to manipulate.
However, it is not easy to identify repertoires of nanobodies with sufficient high affinity to specific targets – current methods have proved too time consuming and difficult – and so many researchers continue to use antibodies. Now, that might change, thanks to a new technique published in Nature Methods and led by The Rockefeller University in New York, NY, as co-author Michael Rout, professor and head of Rockefeller’s Laboratory of Cellular and Structural Biology, explains:
“Nanobodies have tremendous potential as versatile and accessible alternatives to conventional antibodies, but unfortunately current techniques present a bottleneck to meeting the demand for them. We hope that our system will make high-affinity nanobodies more available, and open up many new possible uses for them.”
First, the team made antibodies with high-affinity – that is highly tuned to bind precisely to their molecular targets – and targeted them to find two fluorescent proteins: GFP and mCherry. Biologists use these fluorescent proteins to visualize activity inside cells. Like conventional ways of making antibodies, their technique uses animals at first. In this case, the team started with llamas, which are known to make antibodies that are easily modified to make nanobodies. They immunized the llamas with the two proteins, so their immune systems readily produced the required antibodies.
The next step was crucial in speeding up the production of nanobodies: how to rapidly sequence the genetic code of the high-affinity antibodies – the ones that had the greatest ability to find and bind to the proteins.
The team started by making sequence databases from RNA they found in the antibody-making cells of the immunized llamas’ bone marrow. Then, using blood samples from the same llamas, they selected the antibodies most tightly bound to the target proteins and chemically cut them into smaller sections. For making the nanobodies, they kept only those sections of the antibodies that were tightly bound to the proteins.
Using mass spectrometry and a computer algorithm they called “llama magic,” the team then determined the partial amino acid sequences of the building blocks of the nanobodies, and matched the highest affinity ones with the original RNA sequences they found in the antibody-producing cells.
They then used the antibody cell RNA sequences that matched the sequences of the high-affinity nanobodies to engineer bacteria to mass produce the nanobodies. New technique generates much larger repertoire of high-affinity nanobodies. The next step was to test the new nanobodies. Scientists often use antibodies to isolate a particular part within a cell so they can remove it and study its structure. So this is what the team did with their new nanobodies. They purified various cell structures tagged with GFP or mCherry, and visualized them in place.
Using their new technique, the team generated 25 types of nanobodies with high affinity for GFP and six for mCherry. This is a much larger repertoire than ones typically produced with conventional methods. A large repertoire is important because it gives researchers more options: they can choose the best nanobodies, eliminate ones that might react with other molecules as well as the target ones, and they can also string two together and attack two places on the same target molecule.
“Given that we can now readily identify suites of high-affinity nanobodies, the future for them as research tools, diagnostics and therapeutics looks bright. In 2012, Medical News Today learned how researchers from the Institute of Tropical Medicine, in Antwerp, Belgium, are working on a way to use “trojan horse” bacteria to release nanobodies to conquer sleeping sickness,Prof. Rout said.
Be a part of Elets Collaborative Initiatives. Join Us for Upcoming Events and explore business opportunities. Like us on Facebook , connect with us on LinkedIn and follow us on Twitter , Instagram.
"Exciting news! Elets technomedia is now on WhatsApp Channels Subscribe today by clicking the link and stay updated with the latest insights!" Click here!




















